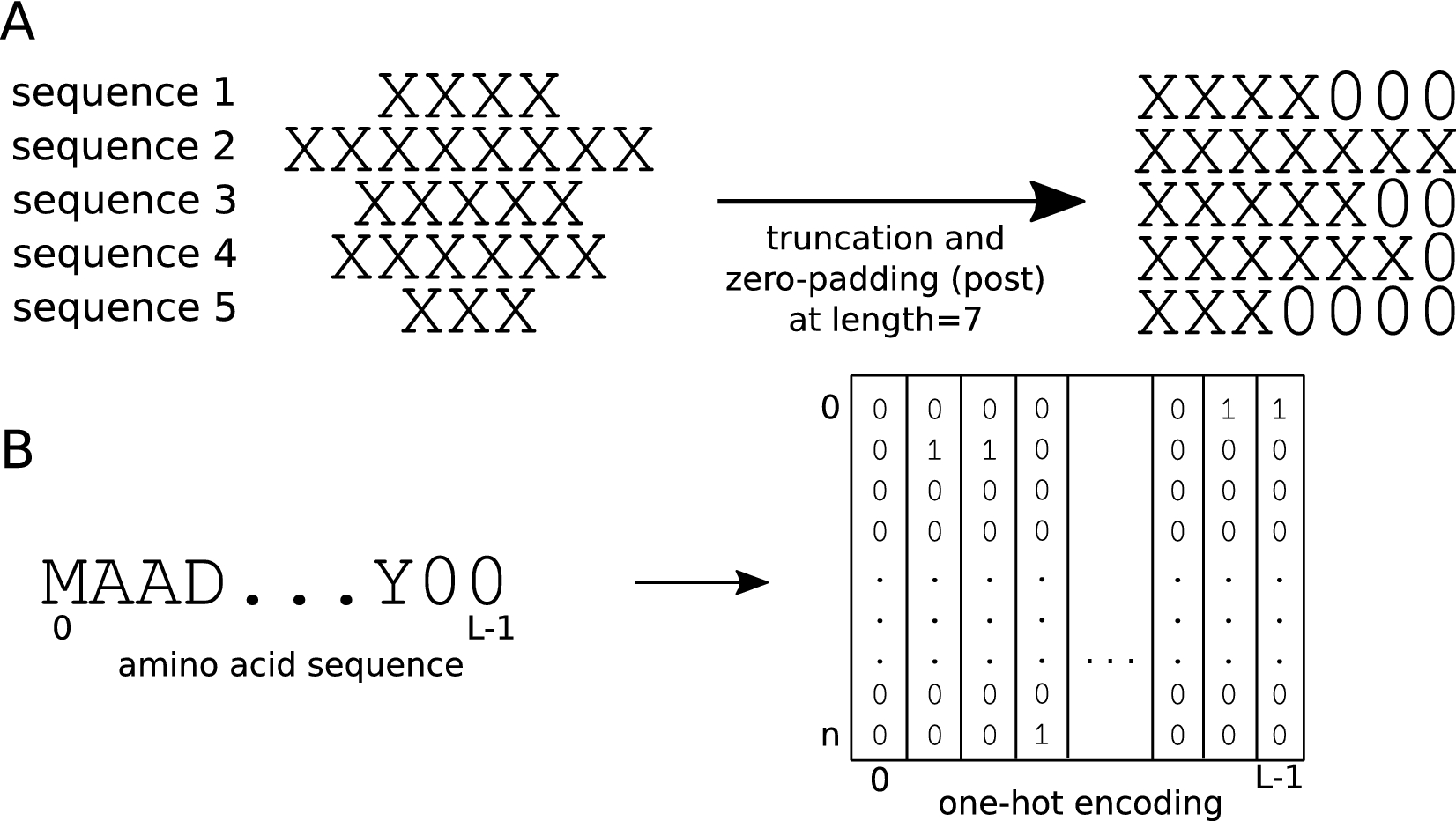
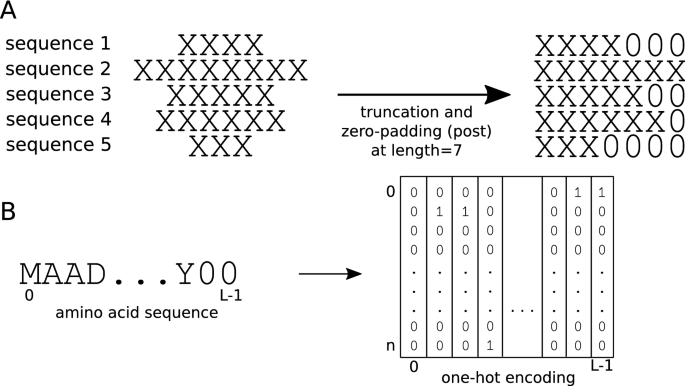
Effect of sequence padding on the performance of deep learning models in archaeal protein functional prediction

Deep learning program to predict protein functions based on sequence information - ScienceDirect

Ángela López del Río

Materials, Free Full-Text

Amino Acid Encoding Methods for Protein Sequences: A Comprehensive Review and Assessment

PDF) Effect of sequence padding on the performance of deep learning models in archaeal protein functional prediction

Summary of Phylogenetic, Functional, Metabolite and Domain

FFP: joint Fast Fourier transform and fractal dimension in amino acid property-aware phylogenetic analysis, BMC Bioinformatics

MECE: a method for enhancing the catalytic efficiency of glycoside hydrolase based on deep neural networks and molecular evolution - ScienceDirect

Deep learning for protein secondary structure prediction: Pre and post-AlphaFold - ScienceDirect

Ángela López del Río

Battery degradation prediction against uncertain future conditions with recurrent neural network enabled deep learning - ScienceDirect

Deep Learning in Protein Structural Modeling and Design - ScienceDirect

Angela LOPEZ-DEL RIO, Universitat Politècnica de Catalunya, Barcelona, UPC, CREB - Biomedical Engineering Research Centre

Deep learning prediction of enzyme optimum pH