Efficient phasing and imputation of low-coverage sequencing data
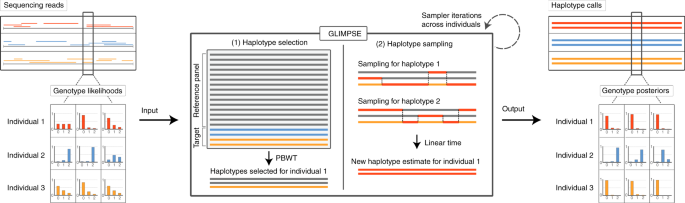

Assessment of the performance of different imputation methods for

PDF) Assessment of the performance of different imputation methods for low-coverage sequencing in Holstein cattle
Discovery and false-positive rates of QCALL for 400 samples with 4.03

Frontiers Ultra Low-Coverage Whole-Genome Sequencing as an
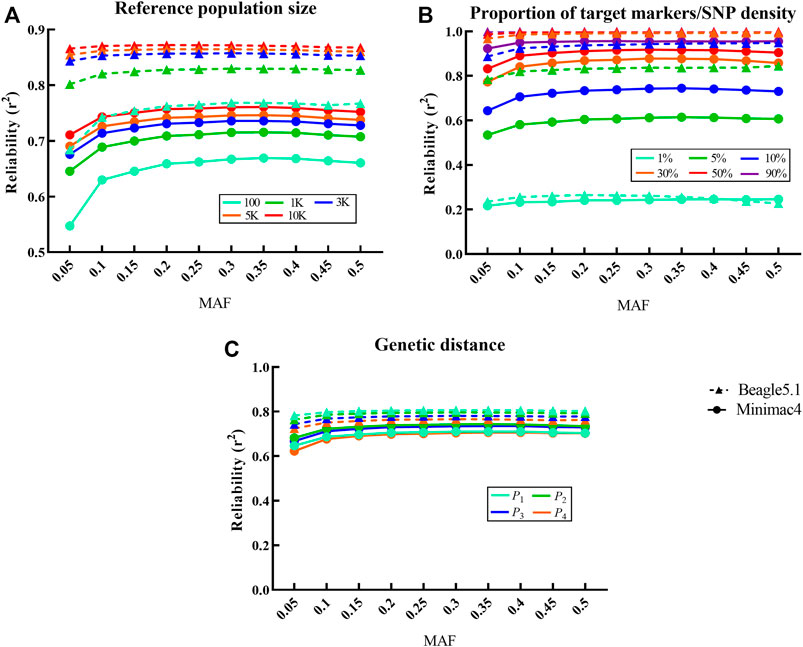
Frontiers Comparison of Genotype Imputation for SNP Array and Low-Coverage Whole-Genome Sequencing Data
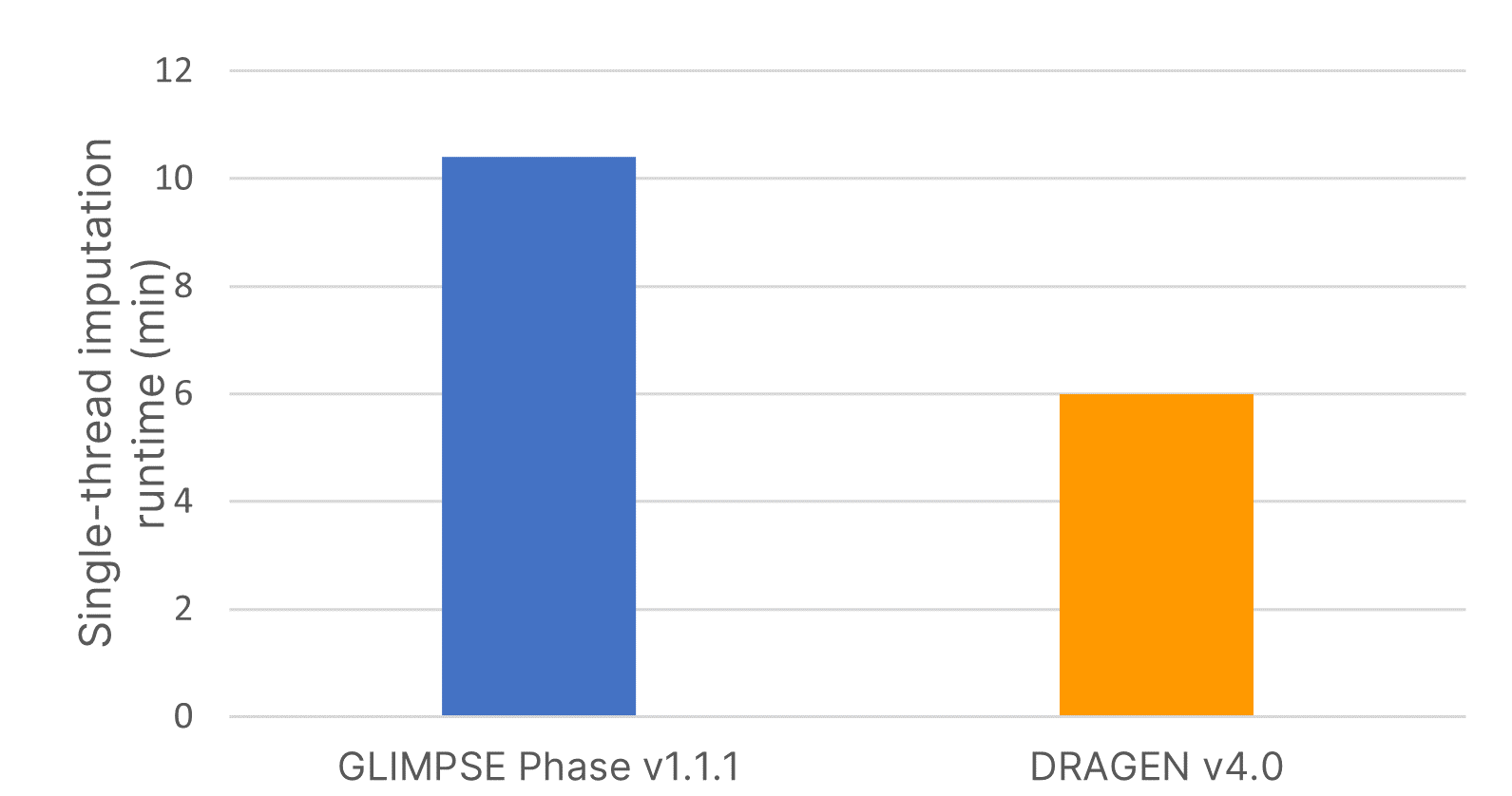
Boosting variant calling performance using a high-quality
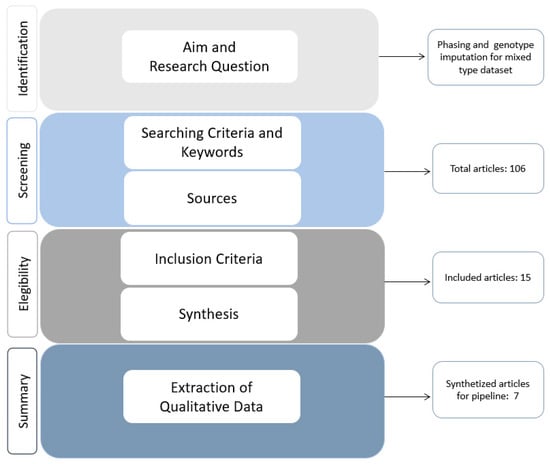
Life, Free Full-Text

E07.1 - Efficient phasing and imputation of low coverage

Imputation of ancient human genomes
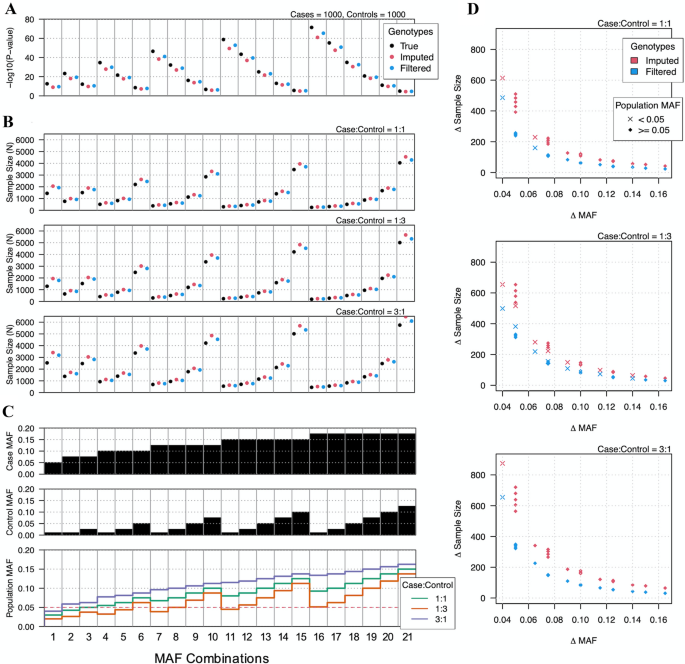
Best practices for analyzing imputed genotypes from low-pass sequencing in dogs

Accurate rare variant phasing of whole-genome and whole-exome sequencing data in the UK Biobank - Abstract - Europe PMC







